Welcome to Iris Biotech
For better service please confirm your country and language we detected.

For better service please confirm your country and language we detected.

Thank you very much for your interest in our products. All prices listed on our website are ex-works, Germany, and may attract customs duties when imported.
You may/will be contacted by the shipping company for additional documentation that may be required by the US Customs for clearance.
We offer you the convenience of buying through a local partner, Peptide Solutions LLC who can import the shipment as well as prepay the customs duties and brokerage on your behalf and provide the convenience of a domestic sale.
Continue to Iris Biotech GmbHSend request to US distributorPublished on 30/05/2023

Fluorogenic substrates are the method of choice for many assays to determine protease content or to measure enzyme kinetics. Compared to absorption measurements, where there is typically only a gradual change in the signal during the course of the reaction, fluorescence offers possibilities to monitor signal changes from (almost) zero emission to bright fluorescence or vice versa. This significantly enhances the signal-to-noise-ration of the experiment and thus the evaluability of the data.
To enable this significant change in fluorescence emission from “dark” to “bright”, there are two basic principles available: Either the protease reaction chemically changes the fluorophore itself and thus its fluorescence behavior, or the protease action changes the spatial distance between a fluorescence donor and a FRET acceptor or quencher.
1) Protease reaction changes the fluorophore – Rhodamine substrates
The first principle is applied for many decades. Coumarin derivatives such as AMC (7-Amido-4-methylcoumarin) are non-fluorescent as long as they are bound to a peptide by an amide bond but start to fluoresce once they are released from the peptide by a protease. However, AMC’s excitation and emission wavelengths are in the UV range and thus the performance of experiments with biological samples is often limited by the presence of UV absorbing biomolecules such as nucleic acids, ribosomes, or many other UV absorbing particles.
Rhodamines such as Rhodamine 110 (Rh 110) have a ca. 100 nm redshift compared to AMC, thus acting far away from the absorption ranges of most biomolecules, enabling a better and undisturbed detection. Their emitted light around 530 nm is yellow to green.
Rh 110 has two free amine groups that can be acylated with amino acids or peptides. As long as both amines are acylated, the Rhodamie’s fluorescence is almost zero. After hydrolysis of the first modification, the fluorescence increases by a factor of ~3500. Subsequent hydrolysis of the second modification gives another fluorescence increase by a factor of ~10.
Typically, and for practical chemical reasons, Rh 110 is acylated symmetrically twice with the same substrate peptide. This mechanism with two subsequent hydrolyses makes the kinetic evaluation of the enzyme kinetics challenging and difficult.
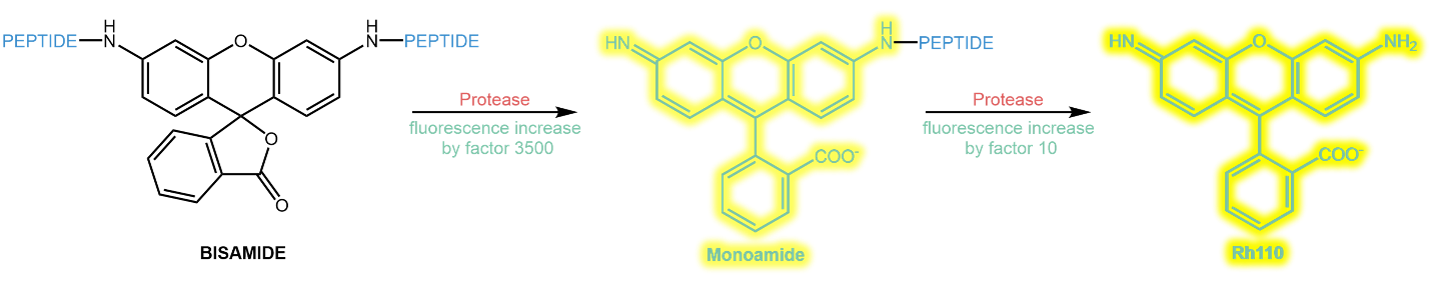
Sequential hydrolysis of a symmetrically substituted Rhodamine bisamide.
With a sophisticated synthetic approach, we can generate asymmetric Rh 110 peptide substrates that only carry one hydrolysable peptide moiety, whereas the second modification spot remains blocked by a non-protease labile modification. This results in a one-step hydrolysis mechanism with high signal-to-noise-ratio that can easily be evaluated:

Single hydrolysis of a blocked Rhodamine bisamide bearing only one hydrolysable peptide moiety.
2) Protease reaction changes the fluorophore’s surrounding
In a combined system of fluorescence donor and acceptor (“FRET pair”) or fluorophore and quencher, the action of the protease changes the spatial distance between the two interacting fluorescence partners.
If the combination in a peptide substrate is, for example, EDANS (= fluorophore) and Dabcyl (= quencher), EDANS will exhibit only weak fluorescence emission as long as Dabcyl is fixed in vicinity. As soon as a protease cuts the connection between the partners, EDANS fluorescence will shine bright:
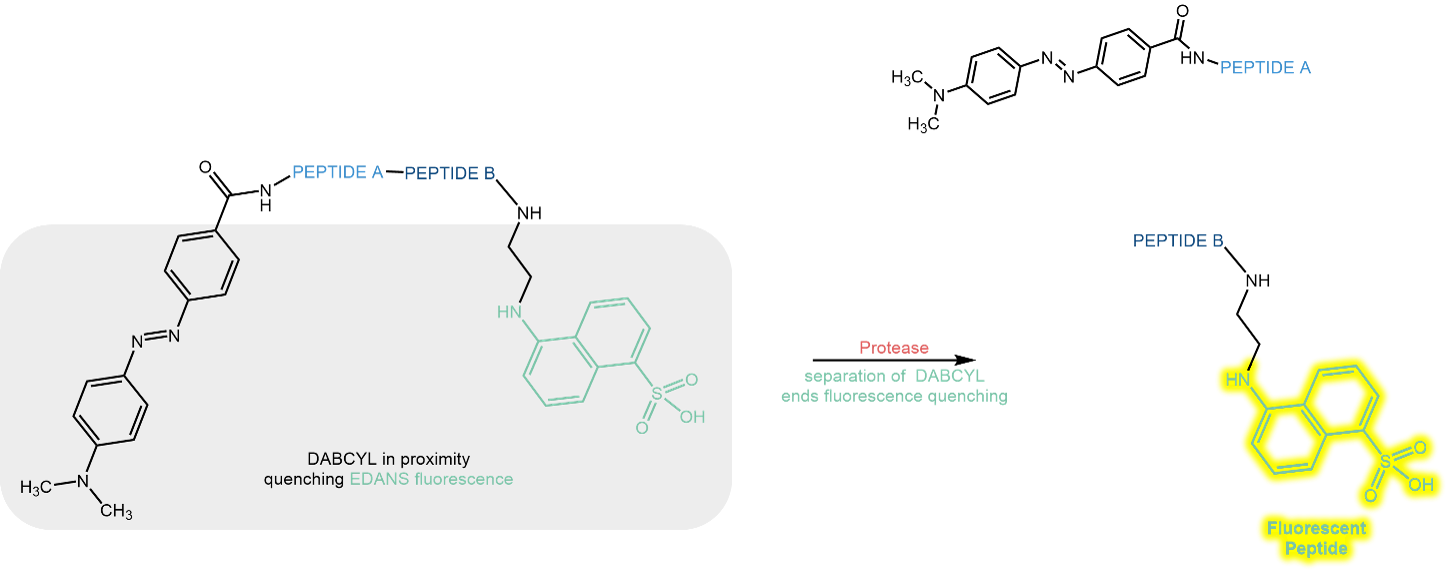
Schematic illustration of the FRET pair EDANS (fluorophore) and Dabcyl (quencher).
Iris Biotech can supply you with several fluorogenic peptide substrates applying one of the described methods to improve and facilitate hydrolytic enzyme detection and characterization. Further to our catalogue substrates for Caspase, Paracaspase (MALT1), Trypsin, Prostatin, Matriptase, or Calpain we can synthesize for you almost any peptide based fluorogenic substrate.
Two of our fluorogenic peptide substrates have recently gained importance in diagnostic and research of SARS-CoV-2. We provide two fluorogenic substrates for the SARS-CoV-2 main protease Mpro (3CLpro), which is one of the major targets in the search for pharmaceuticals to inhibit the virus multiplication. One is using an asymmetric Rh110 (LS-4190), whereas the other makes use of the EDANS / Dabcyl quenching pair (LS-4180).
Don’t hesitate to contact us for an optimized fluorogenic peptide substrate for your application!
References:
Screening a Library of FDA-Approved and Bioactive Compounds for Antiviral Activity against SARS-CoV-2; S. B. Biering, E. Van Dis, E. Wehri, L. H. Yamashiro, X. Nguyenla, C. Dugast-Darzacq, T. G. W. Graham, J. R. Stroumza, G. R. Golovkine, A. W. Roberts, D. M. Fines, J. N. Spradlin, C. C. Ward, T. Bajaj, D. Dovala, U. Schulze-Gamen, R. Bajaj, D. M. Fox, M. Ott, N. Murthy, D. K. Nomura, J. Schaletzky, S. A. Stanley; ACS Infect. Dis. 2021; 7(8): 2337–2351. https://pubs.acs.org/doi/full/10.1021/acsinfecdis.1c00017
The SARS-CoV-2 main protease as drug target; S. Ullrich, C. Nitsche; Bioorg Med Chem Lett. 2020; 30: 127377; https://doi.org/10.1016/j.bmcl.2020.127377
Structural basis for the inhibition of SARS-CoV-2 main protease by antineoplastic drug carmofur; Z. Jin, Y. Zhao, Y. Sun, B. Zhang, H. Wang, Y. Wu, Y. Zhu, C. Zhu, T. Hu, X. Du, Y. Duan, J. Yu, X. Yang, X. Yang, K. Yang, X. Liu, L. W. Guddat, G. Xiao, L. Zhang, H. Yang, Z. Rao; Nat. Struct. Mol. Biol. 2020; 27: 529-532; https://doi.org/10.1038/s41594-020-0440-6
Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved a-ketoamide inhibitors; L. Zhang, D. Lin, X. Sun, U. Curth, C. Drosten, L. Sauerhering, S. Becker, K. Rox, R. Hilgenfeld; Science 2020; 368: 409-412; https://doi.org/10.1126/science.abb3405
Structural Determinants of MALT1 Protease Activity; C. Wiesmann, L. Leder, J. Blank, A. Bernardi, S. Melkko, A. Decock, A. D‘Arcy, F. Villard, P. Erbel, N. Hughes, F. Freuler, R. Nikolay, J. Alves, F. Bornancin, M. Renatus; J. Mol. Biol. 2012; 419: 4-21. https://doi.org/10.1016/j.jmb.2012.02.018
Chemical Synthesis of Ubiquitin, Ubiquitin-Based Probes, and Diubiquitin; F. El Oualid, R. Merkx, R. Ekkebus, D. S. Hameed, J. J. Smit, A. de Jong, H. Hilkmann, T. K. Sixma, H. Ovaa; Angew. Chem. Int. Ed. 2010; 49: 10149-10153. https://doi.org/10.1002/anie.201005995
Development of Novel Assays for Proteolytic Enzymes Using Rhodamine-Based Fluorogenic Substrates; S. K. Grant, J. G. Sklar, R. T. Cummings; J. Biomol. Screen. 2002; 7: 531-540. https://doi.org/10.1177/1087057102238627
A sensitive fluorescence intensity assay for deubiquitinating proteases using ubiquitin-rhodamine110-glycine as substrate; U. Hassiepen, U. Eidho!, G. Meder, J.-F. Bulber, A. Hein, U. Bodendorf, E. Lorthiois, B. Martoglio; Anal. Biochem. 2007; 371: 201-207. https://doi.org/10.1016/j.ab.2007.07.034
Rhodamine-based compounds as fluorogenic substrates for serine proteinases; S. P. Leytus, L. L. Melhado, W. F. Mangel; Biochem. J. 1983; 209: 299-307. https://doi.org/10.1042/bj2090299